LAM-PCR HTGTS from Cas9:RAG1A on-target and off-target bait sites (a)... | Download Scientific Diagram
Outils - CRIBL - Contrôle de la Réponse Immune B et Lymphoproliférations UMR CNRS 7276 / INSERM U 1262

WO2016081798A1 - Methods relating to the detection of recurrent and non-specific double strand breaks in the genome - Google Patents

CTCF-Binding Elements Mediate Accessibility of RAG Substrates During Chromatin Scanning. - Abstract - Europe PMC
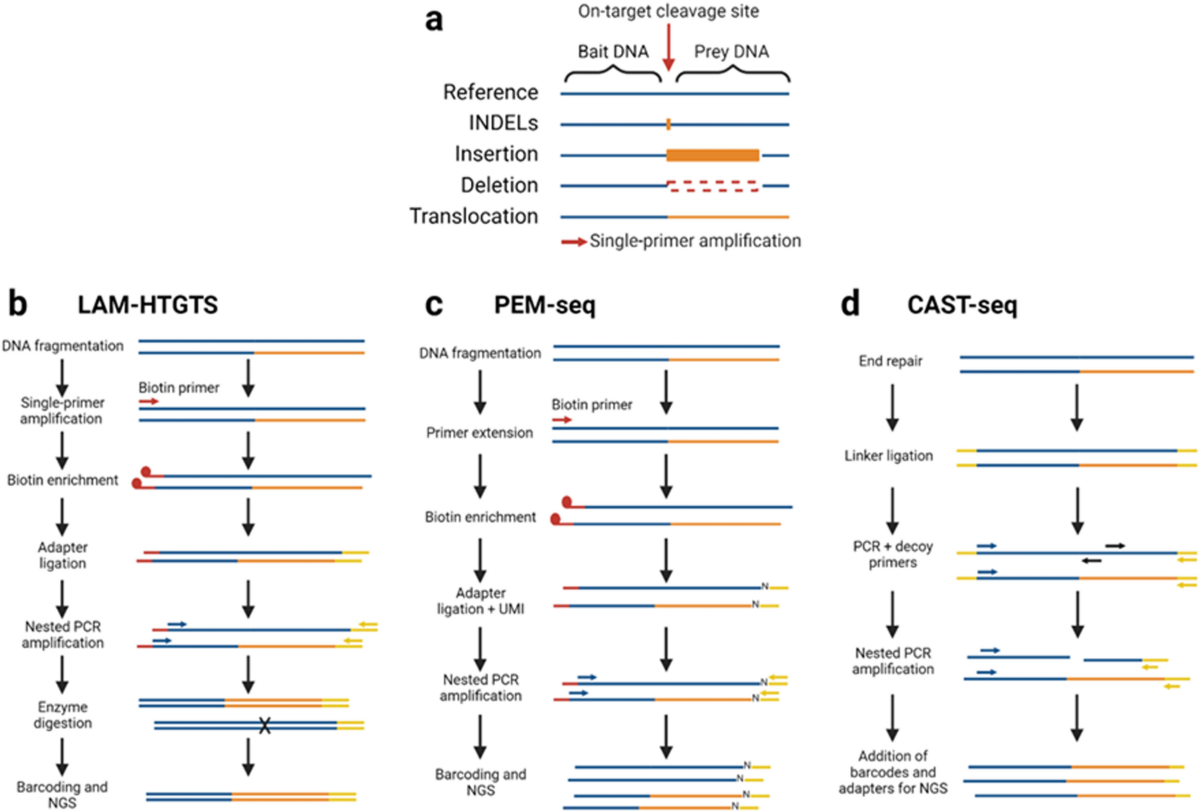
Unintended CRISPR-Cas9 editing outcomes: a review of the detection and prevalence of structural variants generated by gene-editing in human cells | Human Genetics
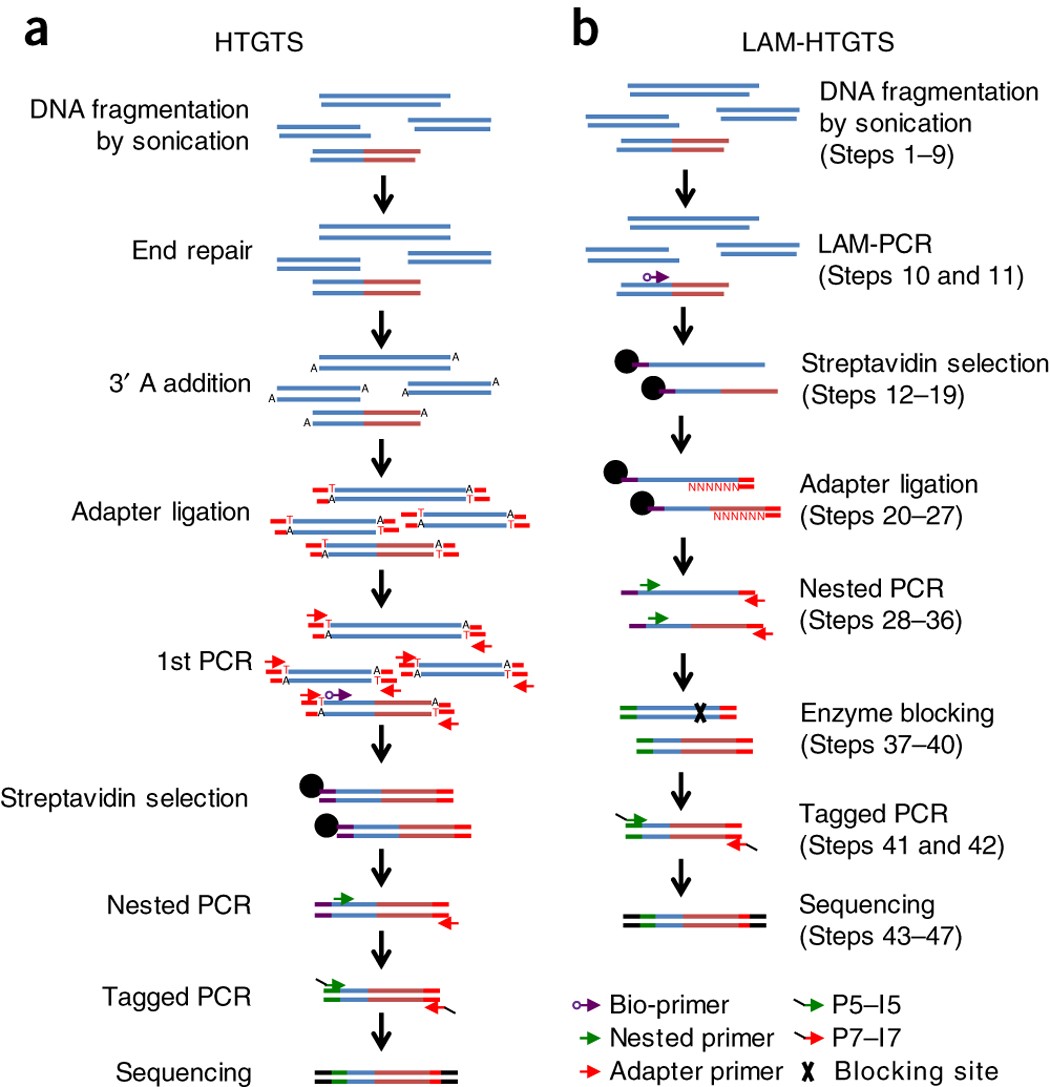
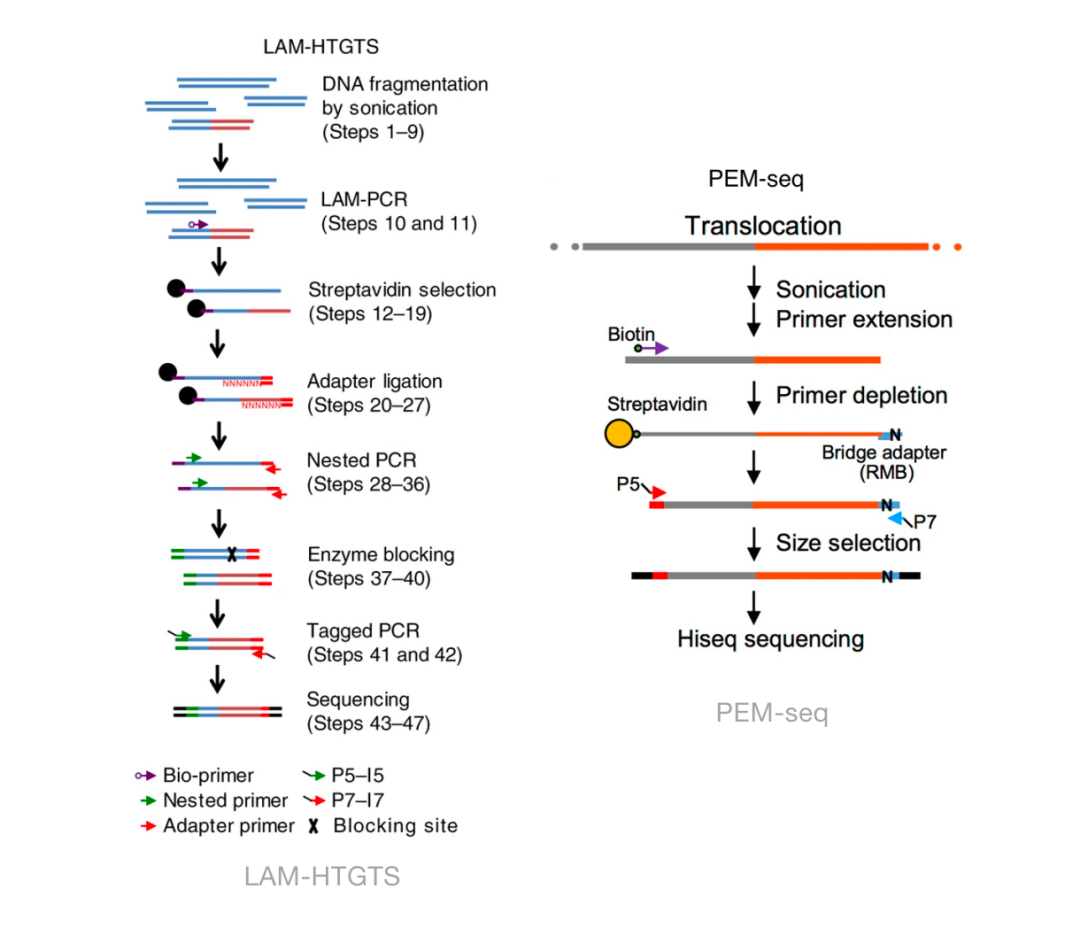
Detecting DNA double-stranded breaks in mammalian genomes by linear amplification–mediated high-throughput genome-wide translocation sequencing | Nature Protocols
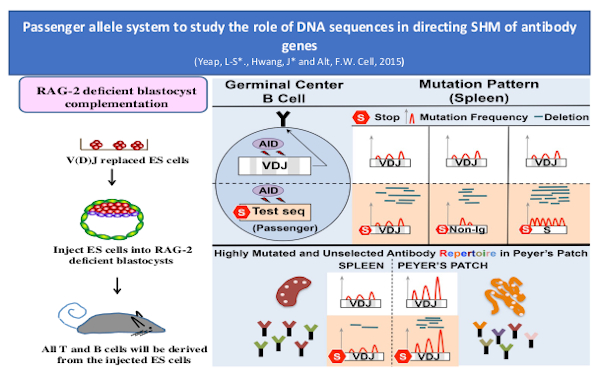
Recent & Ongoing Studies | Research Projects | Alt Laboratory Research | Research Labs | Research | Boston Children's Hospital

Detecting DNA double-stranded breaks in mammalian genomes by linear amplification-mediated high-throughput genome-wide translocation sequencing. - Abstract - Europe PMC

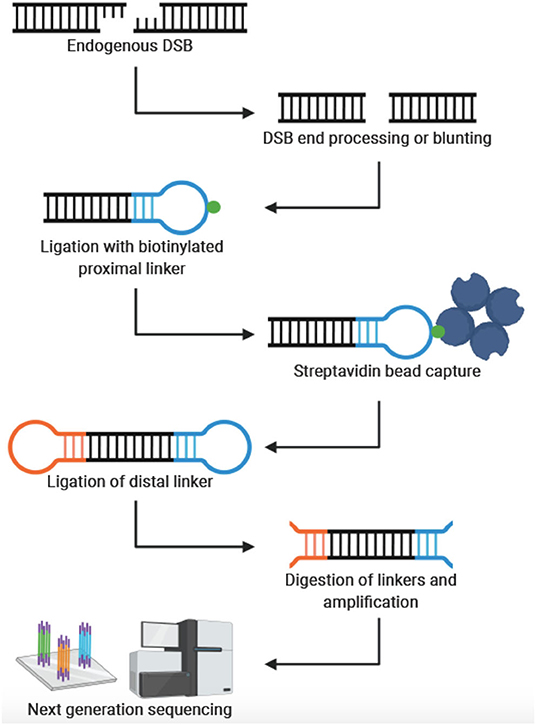
Genes | Free Full-Text | Endogenous DNA Double-Strand Breaks during DNA Transactions: Emerging Insights and Methods for Genome-Wide Profiling

RAG Chromatin Scanning During V(D)J Recombination and Chromatin Loop Extrusion are Related Processes - ScienceDirect

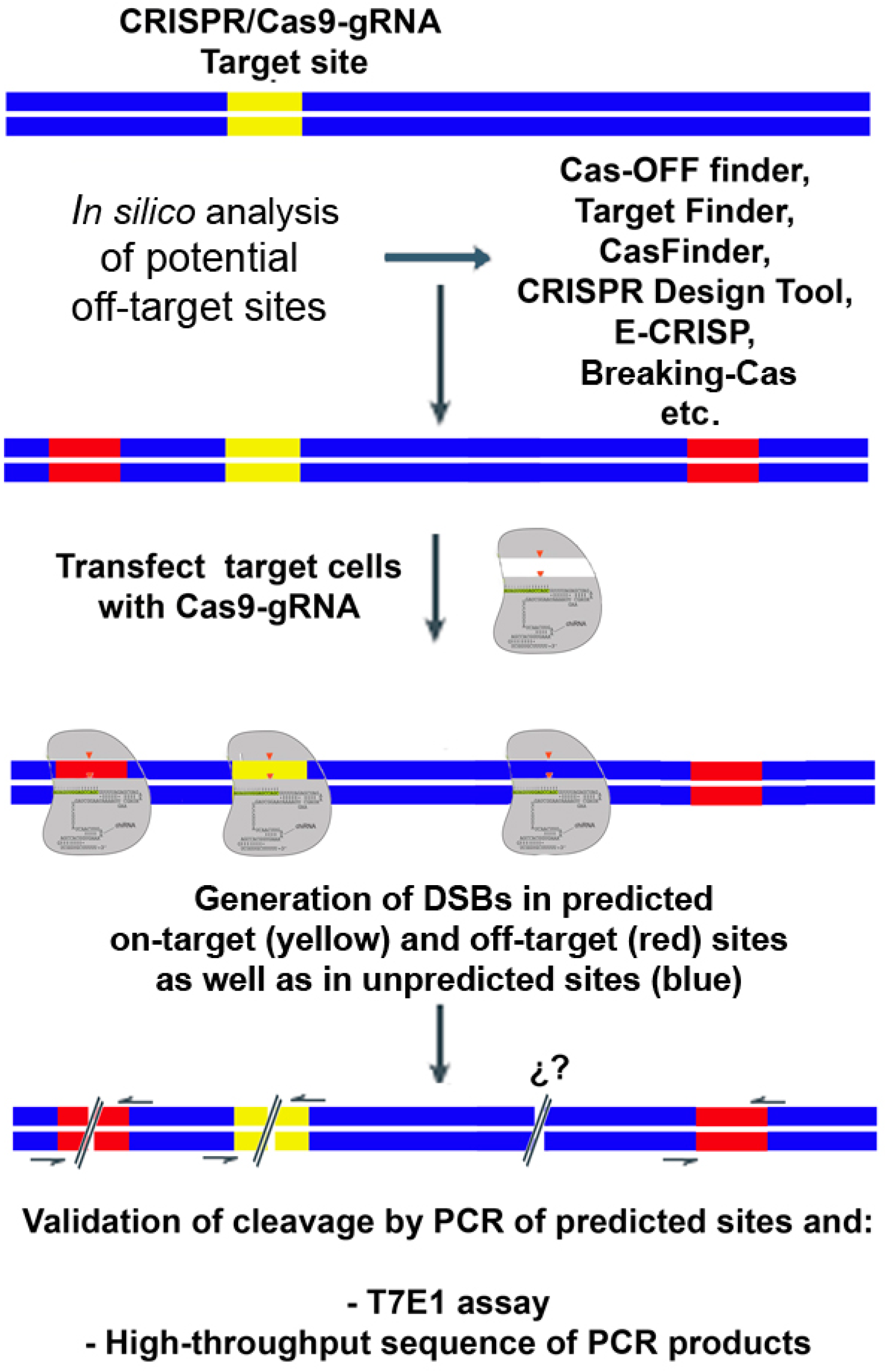
The genome editing revolution: A CRISPR‐Cas TALE off‐target story - Stella - 2016 - BioEssays - Wiley Online Library
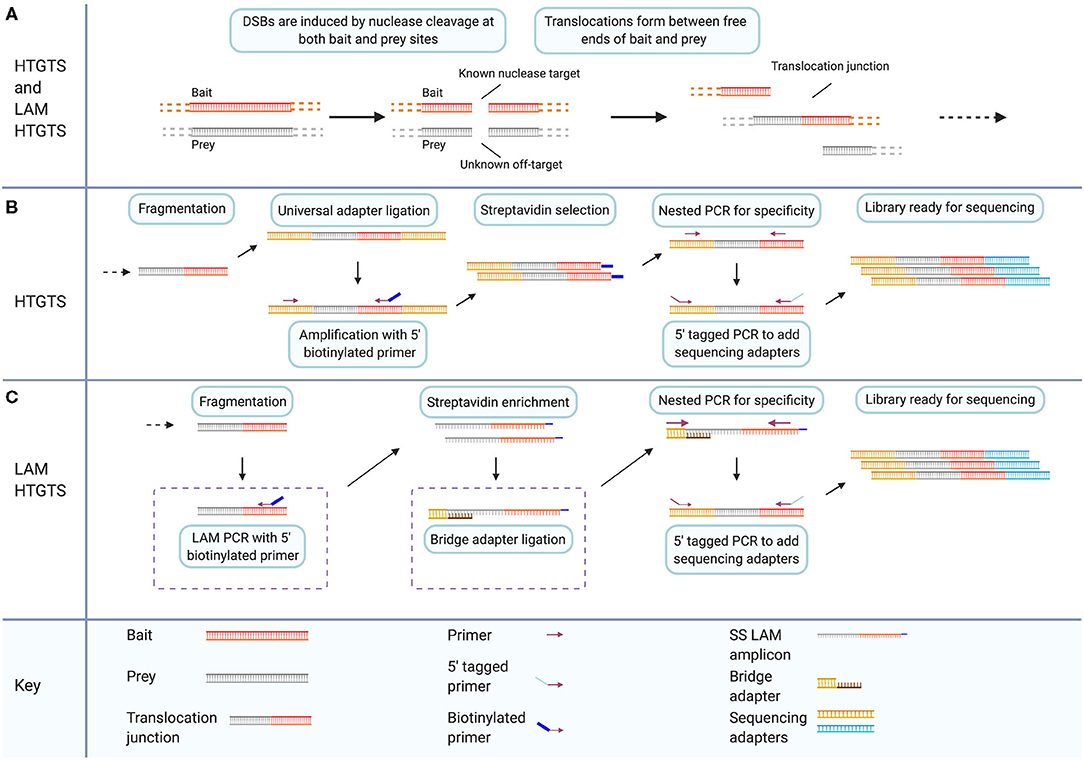
Frontiers | Off-Target Analysis in Gene Editing and Applications for Clinical Translation of CRISPR/Cas9 in HIV-1 Therapy

WO2016081798A1 - Methods relating to the detection of recurrent and non-specific double strand breaks in the genome - Google Patents

Unintended CRISPR-Cas9 editing outcomes: a review of the detection and prevalence of structural variants generated by gene-editing in human cells | Human Genetics

Detecting DNA double-stranded breaks in mammalian genomes by linear amplification–mediated high-throughput genome-wide translocation sequencing | Nature Protocols

LAM-HGTGTS (Linear Amplification-mediated high-throughput genome-wide translocation sequencing) Our Working Protocol.
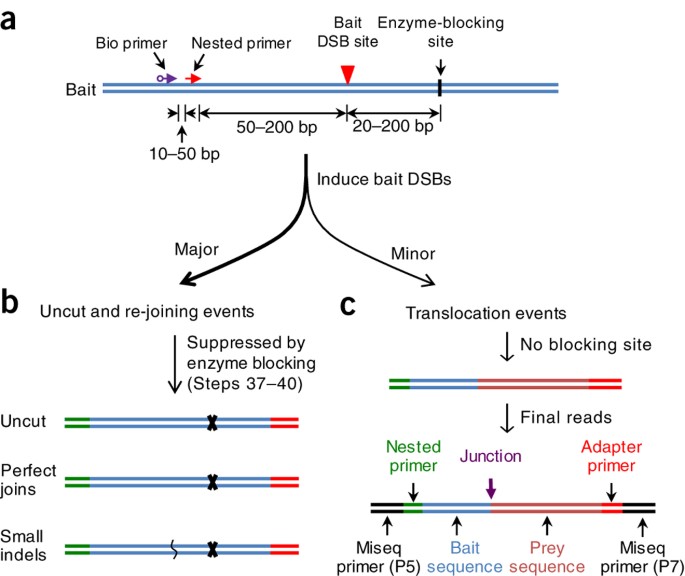
Detecting DNA double-stranded breaks in mammalian genomes by linear amplification–mediated high-throughput genome-wide translocation sequencing | Nature Protocols

Systematic overview on the most widespread techniques for inducing and visualizing the DNA double-strand breaks - ScienceDirect